Ancient DNA Study Reveals Human Evolution Is Happening Faster Than We Thought

New research challenges long-standing assumptions about human evolution, revealing that natural selection has been more active—and more recent—than once believed.
A sweeping analysis of ancient DNA from nearly 16,000 people is reshaping how scientists understand human evolution. By tracking genetic changes across more than 10,000 years in West Eurasia, researchers found that natural selection has been far more active in recent human history than once believed.
For years, evidence for directional selection was surprisingly limited. Only about 21 clear cases had been identified. This process occurs when a specific gene variant provides a survival or reproductive advantage and becomes more common over time, such as the ability to digest milk into adulthood. Because so few examples were known, scientists assumed that this type of evolution played only a minor role after humans spread out of Africa roughly 300,000 years ago.
By analyzing a vastly expanded dataset and applying new statistical tools, researchers uncovered hundreds of gene variants that rose or fell in frequency over time. The findings suggest that human evolution did not slow down in recent millennia. In some ways, it sped up.
Links to Traits and Health
Many of the genetic variants identified in the study are tied to complex traits seen today, including risks for type 2 diabetes and schizophrenia. Exploring how these traits evolved could improve understanding of human biology and disease, and may eventually guide medical research.
At the same time, the authors caution that modern trait definitions do not always apply to ancient populations. For example, measures like household income have no direct equivalent in prehistoric societies, making it difficult to determine why certain variants were originally advantageous.
The study, led by researchers at Harvard University, was published in Nature.
“With these new techniques and large amount of ancient genomic data, we can now watch how selection shaped biology in real time,” said Ali Akbari, the study’s first author. “Instead of searching for the scars natural selection leaves in present-day genomes using simple models and assumptions, we can let the data speak for itself.”
“This work allows us to assign place and time to forces that shaped us,” said senior author David Reich.
10,000 ancient genomes, new computational methods
Since 2010, when scientists first recovered genome-wide data from ancient human remains, the field has transformed our understanding of how populations are related across time and geography.
Still, researchers have struggled to track how natural selection influenced genetic variation over the past 10,000 years, even though DNA from this period is often well preserved.
This study overcame that challenge through two major advances.
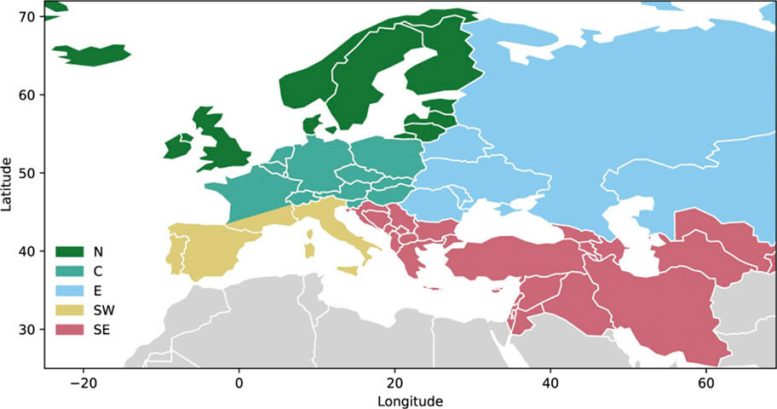
First, Reich’s team spent seven years assembling a large and detailed dataset of ancient DNA from West Eurasia, covering present-day Europe and parts of the Middle East. The effort involved more than 250 archaeologists and anthropologists and produced new genetic data from 10,016 ancient individuals. These were combined with 5,820 previously published ancient genomes and 6,438 modern samples.
“If the goal is to uncover changes in the frequency of genetic variants in the last ten millennia that are greater than can be expected by chance, then we need to detect subtle effects, which requires having thousands of genomes spanning that time period,” Reich explained.
“This single paper doubles the size of the ancient human DNA literature,” he added. “It reflects a focused effort to fill in holes that limited the power of previous studies to detect selection.”
The second advance was a set of computational methods developed by Akbari to separate true signals of directional selection from other influences on gene frequency, such as migration, population mixing, and random fluctuations in small populations.
“Ali developed a powerful technique that could zoom in on the patterns that actually mattered,” said Reich.
Even with these tools, the signal was faint. The researchers estimate that directional selection explains only about 2 percent of all genetic changes observed.
What has natural selection selected for?
That small percentage still represents a significant portion of the genome. The team identified 479 gene variants, or alleles, that were strongly favored or disfavored in West Eurasian populations.
They also traced when and where some of these variants rose or declined. The results show that selection intensified after farming emerged, likely because new diets, environments, and lifestyles created different evolutionary pressures.
More than 60 percent of the selected variants are linked to traits seen in people today, including:
- Light skin tone
- Red hair
- Risk of celiac disease and Crohn’s disease
- Immunity to HIV infection and resistance to leprosy
- Lower chance of male-pattern baldness
- Lower risk of rheumatoid arthritis and alcoholism
- Having the B version of the proteins on red blood cells that confer A, B, and O blood types and influence resistance to infection with bacteria and viruses
In some cases, groups of SNPs were under selection together to influence polygenic traits. Some changes raised the frequency of beneficial traits, including some that are interpreted today as:
- “Health span” traits such as faster walking pace
- Measures of behavioral and social status or cognitive functions, such as scores on intelligence tests, household income, and years of schooling
Other changes reduced the frequency of harmful traits, such as those that are interpreted today as:
- Reduced risk of bipolar disorder and schizophrenia
- Lower body fat percentage, waist-to-hip ratio, and body mass index
- Less susceptibility to tobacco smoking
Some variants rose in frequency and later declined, reflecting shifting environments. For instance, genes linked to susceptibility to tuberculosis and multiple sclerosis showed changing patterns over time.
Not all results are straightforward. One example is a major genetic risk factor for gluten intolerance that became more common after wheat farming began.
Interpreting the Results
The researchers stress that these associations must be interpreted carefully.
A gene’s link to a modern trait does not mean that trait drove its spread in the past. Traits like education level or income did not exist in ancient societies, so they cannot explain why certain variants were favored.
Some variants affect multiple traits, and current databases may not capture their full effects. In other cases, a variant may have increased in frequency simply because it is located near another gene under selection.
It is also possible that some traits influenced by these variants remain unknown today.
Another key point is that these findings are not limited to West Eurasia. The same methods can be applied to other populations with sufficient ancient DNA data to determine which patterns are shared and which are unique.
Reich expects future work to reveal that some selective pressures acted on common traits across different human groups, even as populations spread and adapted to new environments.
What comes next
The researchers have made their data and methods publicly available to support further studies.
One next step is to investigate more than 7,600 additional genetic locations that may represent cases of directional selection. Akbari said these sites have better than a 50 percent chance of “being real examples of directional selection” and deserve closer examination.
Applying the methods to other regions and older time periods is another priority.
“To what extent will we see similar patterns in East Asia or East Africa or Native Americans in Mesoamerica and the central Andes?” Reich asked. “If we can’t use ancient DNA to study the most important period in human evolution 1 million to 2 million years ago, then at least we can study selective pressure on human genomes during more recent periods of change and learn broader principles.”
Further laboratory studies will also be needed to understand how these genetic changes affect health.
The findings could help identify new factors involved in disease, improving risk prediction and treatment. They may also inform gene therapy research. For example, targeting a gene that has been strongly favored by evolution could carry risks.
“You could speculate that if the variant someone wants to knock out was strongly selected for, it’s probably not the best idea,” he said.
Scientists could also use the new methods to study natural selection in other species. Such work could uncover alleles that have made cattle or chickens well-suited to domestication, Akbari suggested, or that have helped animals adapt to changes in climate.
The possibilities are enticing for deepening our appreciation of human diversity, history, and health, Reich said.
“This paper shows how complex selection can be and provides an opportunity to consider the richness of variation in human populations,” he said.
Reference: “Ancient DNA reveals pervasive directional selection across West Eurasia” by Ali Akbari, Annabel Perry, Alison R. Barton, Mohammadreza Kariminejad, Steven Gazal, Zheng Li, Yating Zeng, Alissa Mittnik, Nick Patterson, Matthew Mah, Xiang Zhou, Alkes L. Price, Eric S. Lander, Ron Pinhasi, Nadin Rohland, Swapan Mallick and David Reich, 15 April 2026, Nature.
DOI: 10.1038/s41586-026-10358-1
This research was supported by the John Templeton Foundation (grant 61220), the Allen Discovery Center for Human Brain Evolution (a Paul G. Allen Frontiers Group advised program of the Allen Family Philanthropies), the Howard Hughes Medical Institute, the National Institutes of Health (grant HG012287), a private gift from Jean-François Clin, and the European Research Council (grant 834087, COMMIOS). The research was conducted using the UK Biobank resource under Application 16549. The authors also acknowledge support from the Research Computing Group at HMS.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
Source link

