Your DNA Is Constantly Moving, and It Could Hold the Key to Cancer and Autism

Human DNA constantly refolds in 3D space, and these looping dynamics regulate gene expression and cell identity. Active genes show faster structural turnover, linking genome organization to diseases like cancer and developmental syndromes.
How does the body pack and manage the enormous amount of genetic information required to build a person? And what are the consequences when that system goes wrong? Jesse Dixon, MD, PhD, has dedicated his research to understanding how the genome is arranged in 3D space, knowing that errors in this structure can contribute to cancers and developmental conditions, including autism-related disorders.
His lab’s newest findings strengthen the idea that the genome’s three-dimensional arrangement is constantly shifting. By studying several types of human cells, the team discovered that different parts of the genome unfold and refold at distinct speeds, influencing how genes are regulated and expressed.
Nature Genetics Study Reveals Therapeutic Targets in Genome Folding Machinery
The research, published in Nature Genetics and supported by federal grants and private philanthropy, identifies potential pathways for preventing the harmful folding errors linked to cancer and developmental disease.
“There are six billion base pairs in your genome, and in the last decade we’ve been learning about the molecular machines that fold and organize that massive amount of information,” says Dixon, senior author of the study and associate professor and holder of the Helen McLoraine Developmental Chair at Salk. “What’s interesting is that this folding doesn’t just happen once and then the genome stays put—it seems to be constantly unfolding and refolding. Our study gives us a better idea of where and how often the genome is doing this, which ultimately adds to our understanding of those molecular machines, and, in turn, what may be going on when they dysfunction during cancers or developmental disorders.”
Cohesin Loops and the Mechanics of DNA Organization
Each human cell contains two meters of DNA—an extraordinary length of genetic material that encodes every protein and drives every cellular process. Embedded within this DNA are tens of thousands of genes, short sequences that provide instructions for building proteins or controlling their production.

All of this information must fit inside the cell nucleus while remaining accessible when needed. The genome must be compact, flexible, and capable of bringing distant regions into contact. Cells solve this challenge by forming loops in the DNA. These loops are created by a protein complex called cohesin, assisted by another protein, NIPBL, which helps cohesin travel along DNA and shape these structures.
Earlier research indicated that cohesin-based loops are not permanent. They repeatedly assemble and break apart. Recognizing this constant motion raised new questions. How frequently do loops form and dissolve? Are some genomic regions more dynamic than others? What precise role does NIPBL play in controlling this movement?
Folding Dynamics Influence Gene Expression and Cell Identity
“Current data around the spatial organization of the genome suggest that genome folding has little impact on gene expression—but we thought, perhaps we just aren’t looking at it in the right way,” says first author Tessa Popay, PhD, a postdoctoral researcher in Dixon’s lab. “By specifically disrupting folding dynamics, we were able to identify the aspects of spatial genome organization that contribute to gene regulation and expression.”
To test this idea, the researchers reduced levels of NIPBL in immortalized human retinal pigment epithelial (RPE-1) cells and observed the effects. Without NIPBL, cohesin struggled to move along DNA and form loops. As a result, the genome began to open up. This unfolding did not occur evenly. Some regions relaxed quickly, while others changed gradually over many hours.
The pace of these changes appeared to reflect what the genes in those regions were doing. Loops that remained stable for hours were typically found in silent genome regions where genes were inactive. In contrast, loops that turned over rapidly were linked to expressed genome regions where genes were actively directing cell-type-specific tasks.
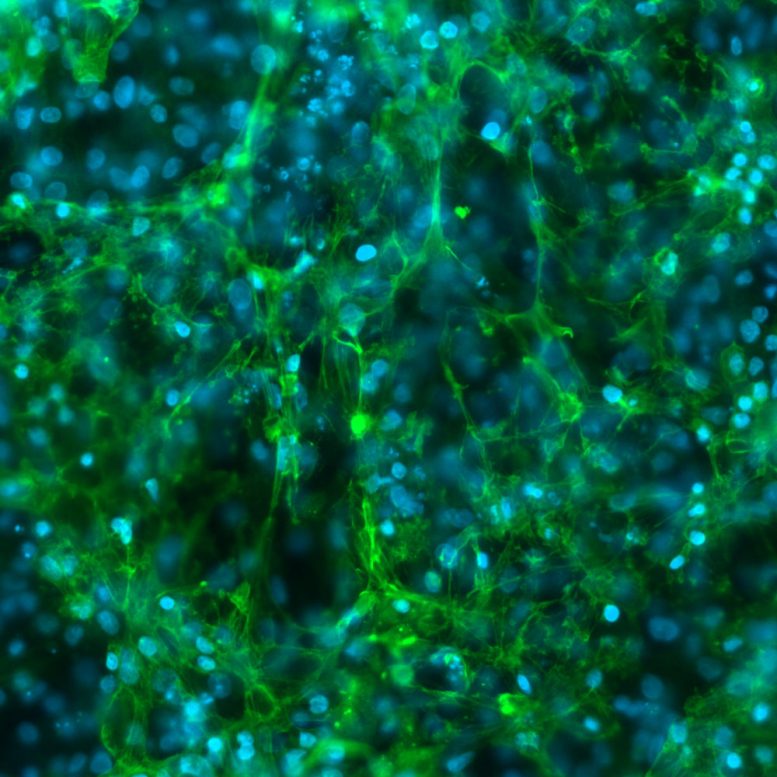
Immortalized retinal pigment epithelial cells (RPE-1) dividing, showcasing the incredible ability for the body to compact, store, duplicate, and repack long strands of genetic information. Credit: Salk Institute
Cell Type–Specific Genome Dynamics and Disease Implications
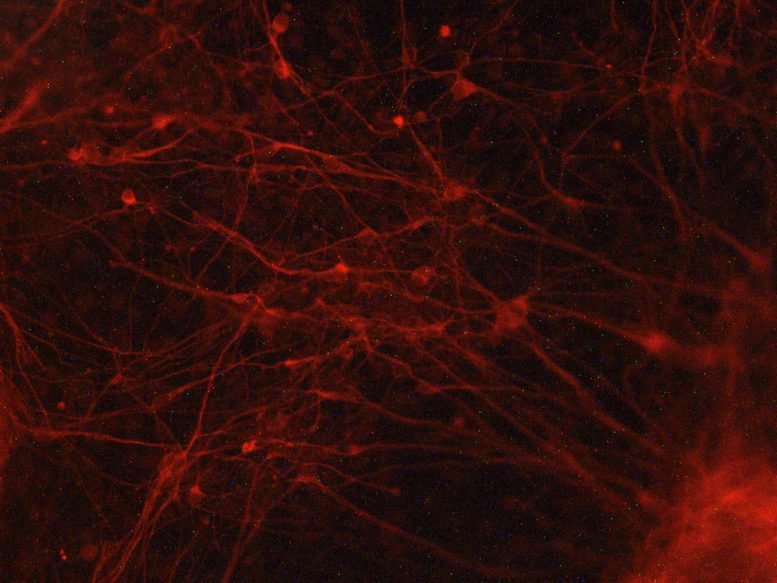
To determine whether these patterns influenced cell identity, the team studied heart cells and neurons generated from human induced pluripotent stem cells (iPSCs). They found that dynamic folding was especially important at genes tied to heart function in heart cells and at genes tied to neuronal function in neurons. Because these specialized genes occupy different positions in the genome, flexible folding may help each cell type maintain its unique program of activity.
“One thing this appears to suggest is that the continuous folding and unfolding of our genome may be particularly important for helping a cell ‘remember’ who it is supposed to be by preserving expression of genes that are unique to different cell types,” says Popay.
The researchers propose that regions associated with cell identity are particularly active because repeated loop formation reinforces the necessary gene connections. In this view, the genome repeatedly reestablishes critical interactions, ensuring that cells continue producing the proteins required for their specialized roles.
Genome Folding Defects Linked to Cancer and Developmental Disorders
Although many questions remain, these findings shed light on why defects in genome folding can have widespread consequences.
“These genome folding machineries tightly control cell identity in every cell, so it actually makes a lot of sense that when we see mutations in them, we get these syndromic conditions like Cornelia de Lange syndrome that impact different parts of the body in different ways,” says Dixon. “And cancer is potentially exploiting that same principle, changing where in the genome these dynamics are more important to manipulate cell identity and encourage uncontrolled growth.”
By confirming that the genome’s shifting 3D structure plays a meaningful role in gene regulation, researchers can better link structural changes to disease. This knowledge opens the door to exploring therapies aimed at correcting harmful folding patterns in cancer and developmental disorders, highlighting the broad impact of understanding how our genetic blueprint is organized.
Reference: “Acute NIPBL depletion reveals in vivo dynamics of loop extrusion and its role in transcription activation” by Tessa M. Popay, Ami Pant, Femke Munting, Melodi Tastemel, Morgan E. Black, Nicholas Haghani and Jesse R. Dixon, 16 February 2026, Nature Genetics.
DOI: 10.1038/s41588-026-02516-y
The work was supported by the National Institutes of Health (U01-CA260700, S10-OD023689, S10-OD034268, P30-CA014195, P30-AG068635, P01-AG073084-04, P30-AG062429), Salk Excellerators Fellowship, Rita Allen Foundation, Pew Charitable Trusts, Howard and Maryam Newman Family Foundation, Helmsley Charitable Trust, Chapman Foundation, Waitt Foundation, American Heart Association Allen Initiative, and California Institute for Regenerative Medicine.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
Source link

